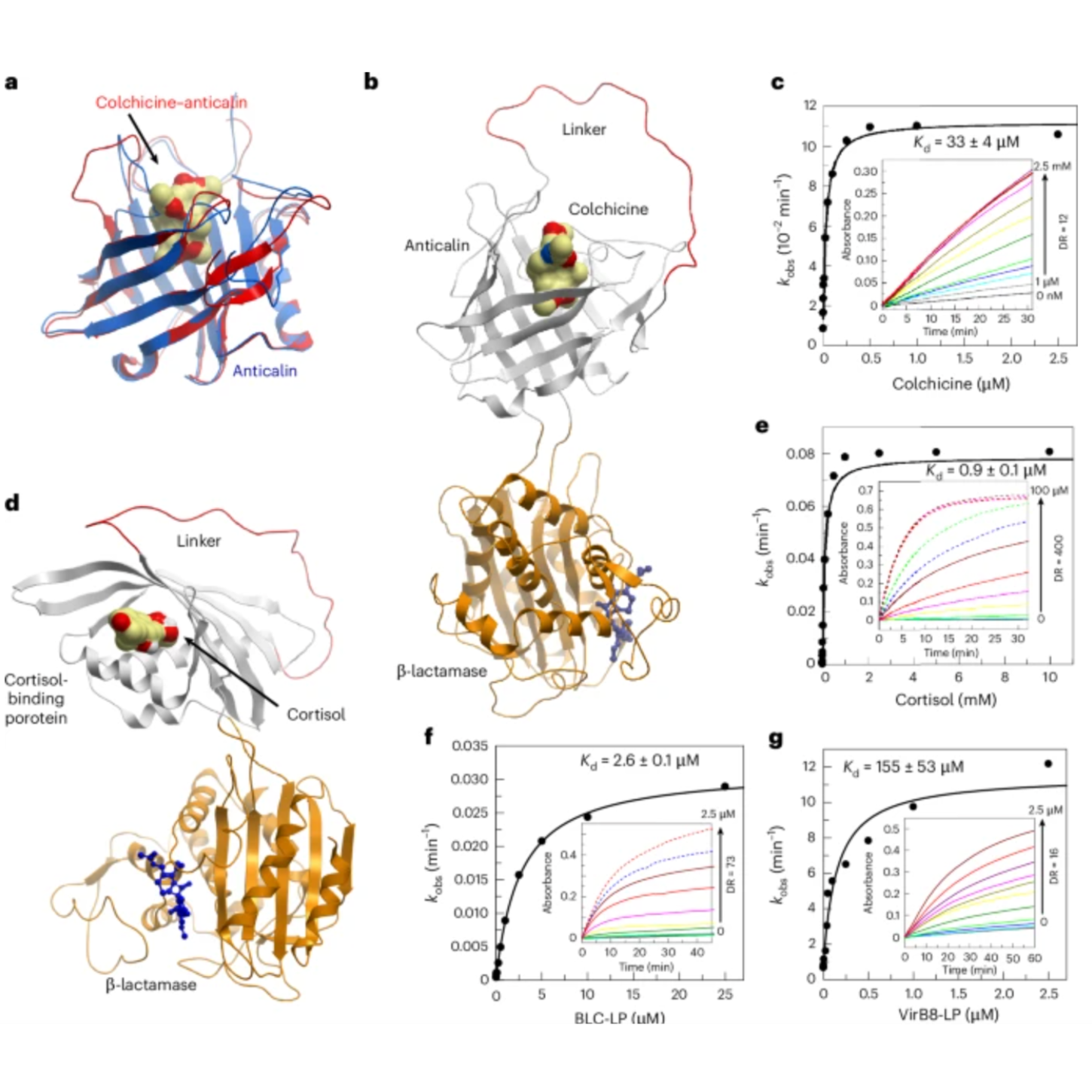
The study demonstrates that machine-learning-designed ligand-binding domains can successfully function as artificial receptors in allosteric protein switches, even without large structural conformational changes. Experimental results show that these engineered proteins can act as versatile biosensors for small molecules, peptides, and proteins, producing measurable outputs such as colorimetric, luminescent, and electrochemical signals. The systems also achieved functional complexity by forming YES and AND logic gates, and showed strong performance in terms of dynamic range, ligand affinity, and catalytic activity across multiple reporter systems.
Mechanistically, the results reveal that ligand binding does not trigger major structural rearrangements but instead reduces conformational entropy, stabilizing the protein and enhancing reporter activity. This was validated using techniques like hydrogen/deuterium exchange mass spectrometry and 19F-NMR (shown in detail in figures on pages 4–5), which demonstrated changes in protein dynamics rather than structure. Additionally, the study highlights that artificial receptors are modular and compatible with different reporter domains, including fully synthetic ones, confirming the generalizability of this design strategy.
Looking ahead, the work opens significant opportunities for synthetic biology and biotechnology. The authors aim to further optimize these systems using machine learning and directed evolution to improve affinity, response time, and stability. Future applications include programmable biological circuits, advanced diagnostics, and real-time biosensing devices. Practical demonstrations—such as engineered E. coli with ligand-dependent survival and bioelectronic sensors for hormone detection (page 7)—indicate strong potential for medical diagnostics, environmental monitoring, and bioelectronic integration.

